How to Choose the Right Protein Expression Systems for Your Research?
In research, selecting the right protein expression systems is crucial for success. Dr. Jane Smith, a leading expert in protein expression, once stated, "The choice of system can be the difference between failure and groundbreaking discoveries." This highlights the importance of careful consideration.
Protein expression systems vary significantly. Each system has unique advantages and limitations. Bacteria, yeast, and mammalian cells all offer differing results. The choice depends on the goals of your research. For instance, bacterial systems produce proteins quickly but often lack post-translational modifications.
Choosing the right system often requires trial and error. Researchers may find themselves stuck between options. Sometimes, initial choices lead to suboptimal yields or incorrect protein folding. This can be frustrating, but it's essential to reflect on what went wrong. The journey is as important as the destination in understanding these complex systems.
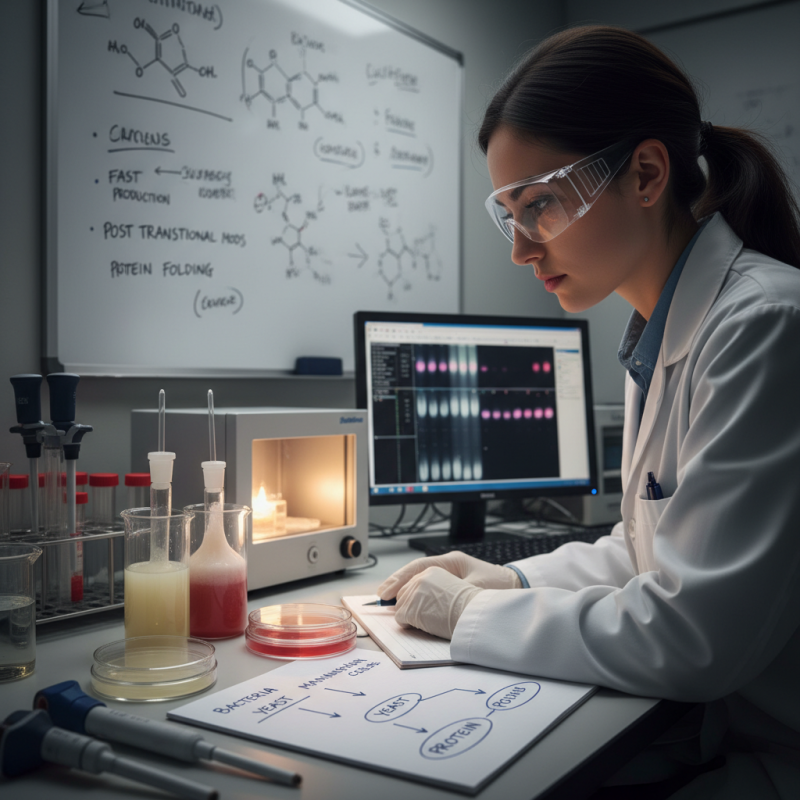

Understanding the Basics of Protein Expression Systems
Protein expression systems are vital tools in molecular biology. These systems enable scientists to produce proteins, which are essential for various research applications. Choosing the right system can be challenging, as each has its strengths and limitations. Understanding the basics of these systems is crucial for informed decisions.
Bacterial expression systems are often the first choice. They are fast and cost-effective. However, they may lack post-translational modifications.
Yeast and insect cell systems offer better modification capabilities but can be more complex. On the other hand, mammalian systems are best for proteins requiring specific folding. Yet, they can be slow and expensive.
Researchers must reflect on their specific needs. Factors like protein size, function, and modifications are key. It is easy to overlook these factors when focusing on speed or cost. Taking the time to assess these requirements may lead to better outcomes in research projects.
Key Factors to Consider When Selecting a Protein Expression System
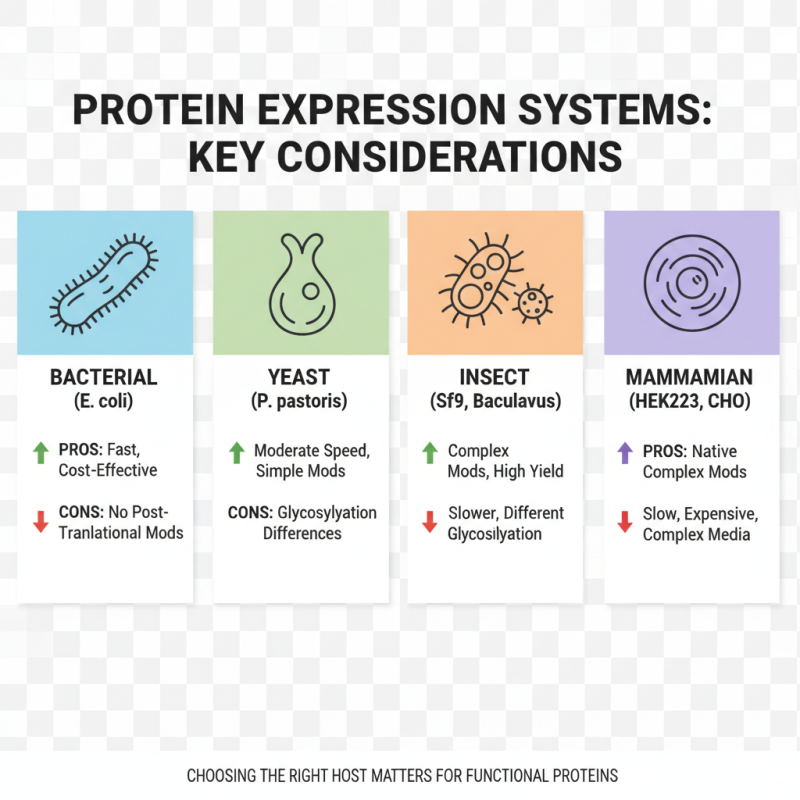

When selecting a protein expression system, several key factors must be considered. The first is the host organism. Each system, whether bacterial, yeast, insect, or mammalian, has its pros and cons. For instance, bacterial systems are often faster and cost-effective but may lack post-translational modifications. In contrast, mammalian cells provide complex modifications necessary for functional proteins but can be slower and more costly.
Another factor is the yield and solubility of the target protein. A recent report indicated that up to 50% of proteins expressed in bacterial systems can form insoluble aggregates. This can obscure results and complicate purification. Yeast systems offer a balance, often achieving higher yields with reasonable solubility. However, understanding the specific requirements of your protein is crucial.
Finally, the scale of expression needed is essential. Many labs prefer systems that can scale up efficiently. A study suggests that mammalian cell systems have a 30% yield on a larger scale compared to bacterial systems. However, scalability requires careful planning and understanding of the system's limitations. Researchers must reflect on their goals and the specific needs of their projects when choosing an expression system.
Common Types of Protein Expression Systems and Their Applications
Choosing the right protein expression system is vital for successful research. Various types exist, each with unique benefits. Bacterial systems, such as E. coli, are commonly used due to low cost and rapid growth. They can perform post-translational modifications but lack complexity. It’s estimated that nearly 50% of recombinant proteins are produced in bacteria, according to recent studies.
Yeast systems, like Saccharomyces cerevisiae, serve as another alternative. They offer more post-translational modifications than bacteria. About 30% of researchers prefer yeast-based systems for eukaryotic protein production. However, challenges in handling intricate pathways can arise.
Insect cell systems also deserve attention. These systems provide a middle ground, achieving better protein folding and post-translational modifications. Data suggests they can boost protein yields by 40% compared to bacterial systems. Still, scaling up can lead to increased costs and time, which may not be ideal for all research goals. Each system has strengths and weaknesses, requiring careful consideration.
Common Types of Protein Expression Systems and Their Applications
Evaluating the Scalability and Yield of Different Systems
When selecting a protein expression system, scalability and yield are crucial factors. Different systems can greatly influence your project’s success. For example, bacterial systems might be fast and economical, yet they often yield proteins that are difficult to purify. Conversely, mammalian systems can produce more complex proteins, but they require more resources and time.
Assess the growth rates of each system. Some microbes double every 30 minutes, while mammalian cells take over 24 hours. This can impact your timeline for research. It's also essential to consider the expected yield. A high yield may sound appealing but may not be achievable in every system. For instance, yeast systems can often produce a substantial amount, but issues with post-translational modifications may arise.
Tips: Start with a small-scale expression test in multiple systems. This can illuminate which option suits your project best. Analyze any inconsistencies in yield carefully. Rethink your choice if results do not align with your expectations. Flexibility in method and mindset is crucial for successful outcomes.
Troubleshooting Common Issues in Protein Expression Processes

Protein expression can be fraught with challenges. Many researchers encounter issues that can hinder their progress. One common problem is low yield. Studies indicate that nearly 30% of protein expression attempts result in less than optimal quantities of target proteins. Factors like plasmid design and host cell choice significantly influence outcomes.
Another issue is protein solubility. Some proteins aggregate, making them difficult to purify. Reports highlight that about 40% of expressed proteins display solubility problems. Utilizing specific fusion tags can enhance solubility. However, this may alter the protein's functionality. It's crucial to balance yield, solubility, and biological activity.
Furthermore, post-translational modifications often pose a challenge in expression systems. Different host cells have varied capabilities to modify proteins. Approximately 20% of proteins fail to fold correctly due to improper modifications. Conducting a thorough troubleshooting process is essential. Adjusting expression conditions, using chaperones, or even switching to a different system may yield better results. A reflective approach is beneficial for overcoming these hurdles.
Related Posts
-
Top 10 Techniques for Protein Expression and Purification Explained
-

Top 10 Insights into Gene Expression You Need to Know?
-

Why Are Recombinant Proteins Important for Biotechnology and Medicine?
-

2026 Top Trends in Life Sciences Innovations and Discoveries?
-
2026 How to Use Human Cell Lines in Research and Development?
-

How to Understand Gene Therapy and Its Impact on Modern Medicine?
